Subprojects
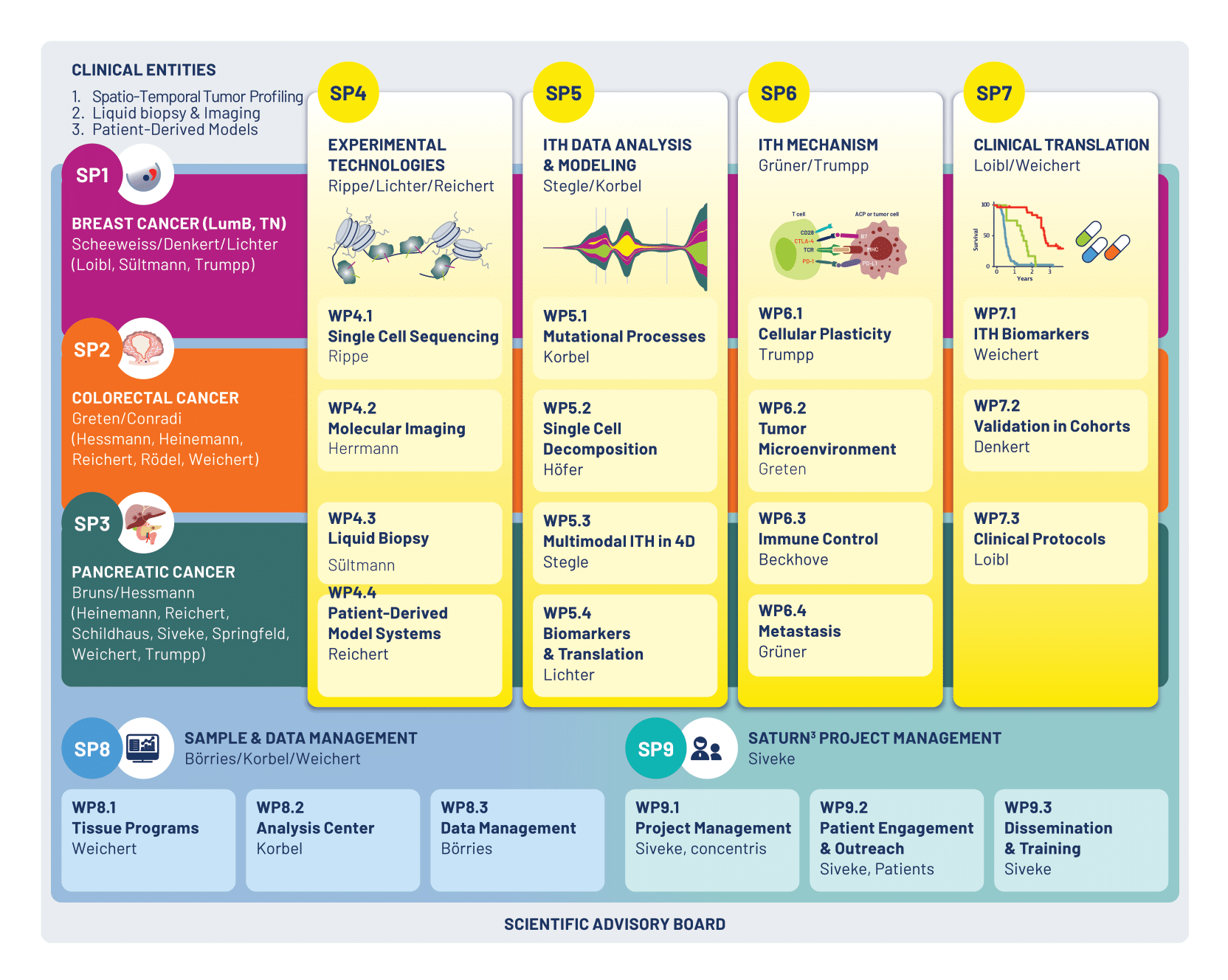
SP1 – Breast cancer
SP Lead: Andreas Schneeweiss, NCT Heidelberg, Heidelberg
SP Co-Lead: Carsten Denkert, Institute of Pathology, Philipps-University Marburg
SP1 will characterize Intratumor Heterogeneity (ITH) in LumB and TNBC in longitudinally collected patient biomaterial of the active precision oncology trials COGNITION (eBC) and CATCH (mBC) as well as paired biobanked samples from phase III trials. ITH in the genome and gene activity landscapes will be determined in tissue and liquid biopsies at single cell (sc) resolution, pre- and post-treatment, with focus on the two clinical scenarios i) chemotherapy -/+ immunotherapy, and ii) molecularly targeted treatment with CDK4/6 and PARP inhibitors. To investigate the role of the microenvironment and cellular plasticity in therapy resistance and metastasis, patient-derived organoids (PDOs) from biopsies and circulating tumor cells (CTCs) will be generated.
The goal is to determine the dynamic spatial and temporal ITH in LumB and TNBC patients to identify therapeutic targets to overcome treatment resistance.
SP2 – Colorectal cancer
SP Lead: Florian Greten, Georg-Speyer-Haus, Frankfurt
SP Co-Lead: Lena-Christin Conradi, Clinic for Surgery, University Medicine Göttingen
SP2 will perform comprehensive longitudinal spatiotemporal molecular analyses of CRC on a single cell level coupled to functional validation employing patient-derived tumor organoids and matched stromal cells to address the following objectives:
- Determine how molecular and cellular heterogeneity impacts on intra-organ and inter-organ differences in metastasis formation and therapy response through longitudinal profiling.
- Examine impact of radiotherapy, conventional chemotherapy and targeted drugs (EGFR, VEGFR) on ITH.
- Functionally validate reciprocal interactions in patient-derived tumor and stromal cells (WP3). Data will be integrated with breast and pancreatic cancer datasets to assess entity overarching ITH principles.
SP3 – Pancreatic cancer
SP Lead: Christiane Bruns, Clinic for Surgery, University Clinic Cologne
SP Co-Lead: Elisabeth Hessmann, Dep. of Gastroenterology, University Medicine Göttingen
SP3 aims to dissect tumor cell-autonomous and context-dependent tumor evolution and subtype-dependent therapy resistance. To this end, and in order to overcome the limitation in sample accessibility evident in metPDAC, five major PDAC centers establish a unique cohort of patients who undergo minimal-invasive multi-site and longitudinal tumor sampling for spatio-temporal molecular analysis. Our profiling strategy is accompanied by lesion-specific patient-derived organoid (PDO) studies in novel assay systems to address cellular plasticity, microenvironment- and epigenetics-related properties. We envision that the elucidation of the therapy-induced spatio-temporal dynamics of the molecular landscapes empowers the refinement of PDAC therapy.
SP3 will pursue the following objectives:
- Dissect spatio-temporal tumoral and microenvironmental evolution at single-cell resolution in metPDAC patients
- Characterize organotropic heterogeneity and epigenomic signatures in existing and novel matched primary and metPDAC biomaterial collections.
- Functionally determine and manipulate PDAC organoid heterogeneity/plasticity. Data will be integrated with breast and colorectal cancer datasets to assess entity overarching ITH principles.
SP4 – Experimental technologies
SP Lead: Karsten Rippe & Peter Lichter, DKFZ, Heidelberg
SP Co-Lead: Maximilian Reichert, University Hospital rechts der Isar (MRI), Munich
SP4, together with the data analysis in SP5, will provide a variety of single cell sequencing (sc-seq) readouts to resolve molecular tumor features in thousands of individual cells and to reveal their evolution during therapy. These in-depth single cell multi-omics analyses of longitudinal sample sets will be combined with PET-CT molecular imaging and liquid biopsies to simultaneously trace development, growth and metastasis in the three tumor entities. Furthermore, complex and advanced patient-derived organoid (PDO) model systems will be generated for investigating mechanisms of tumor heterogeneity and therapy resistance.
SP4 will pursue the following objectives:
- SP4 will acquire integrative sc-seq maps of (epi)genome, transcriptome and cell surface markers in biopsies and PDOs to resolve tumor subclone structure, gene regulation mechanisms and interactions with the tumor microenvironment (TME).
- It conducts molecular imaging to non-invasively visualize tumor microenvironment targets in longitudinal samples.
- SP3 applies liquid biopsy of cell free DNA (cfDNA) to assess therapy response and minimal residual disease (MRD).
- SP3 establishes PDO systems for the functional analysis of heterogeneous tumor phenotypes. The combination of these technologies will generate a comprehensive molecular characterization that defines cell types/states in a given tumor and its TME together with the developmental trajectories during treatment.
- SP4 will provide the technological backbone of Saturn³ to rationalize how tumor heterogeneity and its evolution leads to treatment resistance and to identify key regulators that can be targeted in novel therapeutic approaches.
SP5 – Data analysis & modelling
SP Lead: Oliver Stegle, DKFZ, Heidelberg
SP Co-Lead: Jan Korbel, EMBL, Heidelberg
SP5 will perform state-of-the-art first-line data processing for Saturn³ and develop novel analytical methods including machine learning/AI for dissecting the spatio-temporal dynamics of cancer evolution through integration of bulk, single-cell and spatial omics data. SP5 will also inform key experimental decisions within Saturn³, guiding the selection of the most informative combination of assays and samples to elucidate tumor heterogeneity in depth. SP5 PIs will chair the Analysis and Sample Selection Working Group of Saturn³, which will ensure a deep coordination of analyses, interpretation and experimental design.
SP5 will pursue the following objectives:
- SP5 will pursue state-of-the-art first line processing of diverse data types generated by the consortium, as well as quality control, batch correction, and normalization, to perform genetic, omics and image characterization of tumors.
- It will decompose tumors with clonal resolution, including the temporal integration of clones and alignment of multi-omics profiles to characterize molecular interactions.SP3 applies liquid biopsy of cell free DNA (cfDNA) to assess therapy response and minimal residual disease (MRD).
- SP5 will develop and apply a predictive 4D modelling framework to identify common coordinates of tumor evolution across patients and to deduce mechanistic hypotheses.
- SP5 will identify biomarkers for treatment resistance by performing integrated analyses of our data using a variety of clinical endpoints, including "surrogates" such as data from longitudinal liquid biopsies, and by designing targeted clinically tractable omics assays.
- SP5 will coordinate the feedback of analysis results and findings to other SPs and in particular guide experimental design choices for the consortium, to realize dynamic adjustments to sample and assay design should these become necessary.
SP6 – Mechanistic modeling
SP Lead: Barbara Maria Grüner, West German Cancer Center, University Medicine Essen
SP Co-Lead: Andreas Trumpp, HI-STEM gGmbH, Heidelberg
Utilizing patient-derived organoid (PDO) models derived from serially collected patient samples (primary tumors/progress/relapse/metastases, liquid biopsies) from SP1- 4, and data obtained from their multi-dimensional single cell analysis (SP5), SP6 will decipher the functional and mechanistic impact of intratumor heterogeneity (ITH) on therapy response, resistance development as well as cancer progression and metastasis in patients.
SP6 will address the following specific objectives:
- Delineate and functionally define the cellular programs in cancer cells (i.e. intrinsic) and their surrounding stroma and immune microenvironment (i.e. extrinsic) that underlie cellular plasticity and ITH. Identify the regulatory pathways that determine tumor progression and metastasis.
- Decipher the intrinsic and extrinsic clonal dynamics that cause therapy response and tumor recurrence.
- Define novel therapeutic proof-of-concept approaches disrupting the heterogeneous cancer ecosystems to combat multidrug-resistant tumors and metastasis.
SP7 – Clinical translation
SP Lead: Sibylle Loibl, German Breast Group and Goethe University of Frankfurt, Germany
SP Co-Lead: Katja Steiger, Institute of Pathology, Technical University Munich
SP7 is dedicated to translate the results of SATURN§ into clinical practice and will provide the link between basic research data generated in SP1-6 and the real-world diagnostic and therapeutic situation, paving the way for a full translation of novel tumor heterogeneity concepts into highly individualized everyday patient care. The basis for these extensive validations are large existing biomaterial collections from prospective clinical trials in breast, colorectal and pancreatic cancer already centralized in previous projects. We will transfer new diagnostic ITH markers to standardized assay platforms which can be used for clinical validation in existing cohorts of clinical trials and in the routine diagnostic setting. In addition, we will develop protocols for future clinical trials to prospectively validate the new heterogeneity tests/concepts to generate level I evidence for clinical application and treatment selection.
SP7 will address the following specific objectives:
- Translate the multi-omics and imaging biomarkers generated by the Saturn³ consortium into diagnostically applicable tests.
- Demonstrate that specific approaches testing tumor heterogeneity impact on therapy response and patient outcome (using clinical trial material as well as existing datasets.
- Establish new clinical trial Saturn³ B. Description of Subproject 67 protocols /master protocols for prospective clinical translation to individually target tumor heterogeneity.
SP8 – Sample and data management
SP Lead: Melanie Börries, Institute of Medical Bioinformatics and Systems Medicine (IBSM), Medical Center, University of Freiburg
SP Co-Lead: Jan Korbel, Genome Biology, EMBL, Heidelberg
SP8, will address overarching sample issues and execute a unique image guided postmortem biopsy program (PBP). Within this program, we will generate multisite biopsies from metastatic lesions of BC, CRC and PDAC. Second, an integrated data repository that comprises clinical and molecular data will be established, to pave the way for translating research results into new approaches for diagnosis and treatment of cancer. In pursuing this aim the consortium will consequently adhere to FAIR principles (Findable, Accessible, Interoperable, Reusable) in management of the generated research data and the development of subject-specific and interdisciplinary services, interoperable standards, and interfaces.
SP8 will address the following specific objectives:
- Assembly of a postmortem multisite biopsy cohort of BC, CRC, PDAC and implementation of common tissue processing SOPs
- Provision of integrated clinical datasets with record linkage to imaging and omics data.
- Establishment of a data repository with a user-friendly web portal and presentation of the data using a dashboard. Implementation of an agreed-on, highly interoperable data exploitation strategy that follows the FAIR principles.
SP9 – Project Management
SP Lead: Jens Siveke, West German Cancer Center, University Hospital Essen
SP9 will take care of project management, patient engagement, outreach, dissemination, exploitation, communication, gender issues and training.
SP9 will address the following specific objectives:
- Guarantee an effective project management and governance.
- Monitoring and implementation of risk management and contingency plan.
- Foster patient engagement, outreach to patient communities and the general public.
- Dissemination results and outcome to stakeholders, ensure exploitation of research results, training and sustainability.